CDC algo status
From GlueXWiki
Contents
Add simplifications to offline code to show impact
Using prototype data from CDC_50_50 (run 31942) with offline analysis
50/50 Ar/CO2 and cosmics, 2100V, prototype horizontal
Only 20 samples upsampled
- Event pedestal is mean of 100 samples ending at trigger time
- Find hit threshold crossing
- Upsample 20 samples starting 10 before hit threshold crossing
- Find hit threshold crossing again in upsampled data
- Step back <pedlead> points to find new local pedestal
- Search forward to find high threshold crossing
- Search backward to find low threshold crossing
- Project through both thresholds to find pedestal crossing time
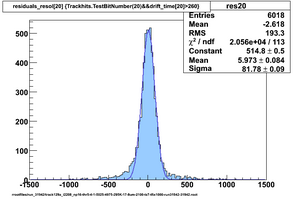
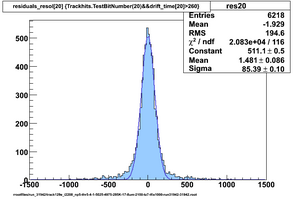
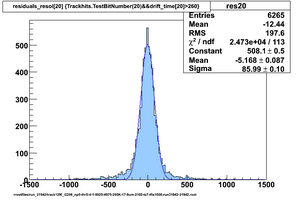
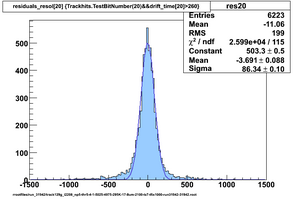
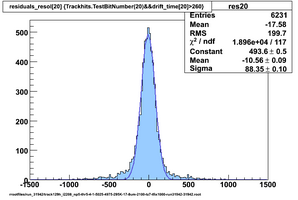
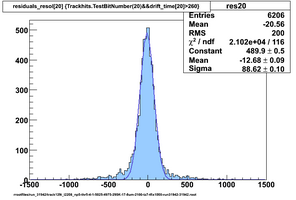
Selected results below
Lower line (c and d) are from single threshold crossings
Implemented in VHDL using ISE...
(d) single (low) threshold crossing without interpolation - takes ~1.4us (includes time for feeding in adc samples), outputs crossing time as integer in units of sample/10. (=0.8ns) (c) single (low) threshold crossing with interpolation - takes ~1.4us (80 ns more than option d), outputs crossing time as integer in units of sample/10. (=0.8ns)