Mattione Update 10312011
From GlueXWiki
Contents
Procedure
- Reconstruct π(1600) -> b1π events with t-slope = 2.0 (was 5.0 for X(2000)) using both the 334 and 1234 segmentations
- Plot Δβ vs p for π+ candidates to see proton, π+ separation.
Details
- The 1234 segmentation is not available in the HDGEANT simulation, so the BCAL shower time for the 1234 segmentation was smeared manually during reconstruction
- 1234: The BCAL hit-time smear was disabled in mc_smear
- 1234: The shower time was smeared by: σsmear = sqrt(σ2t_334 - σ2t_334_NoSmear) * σt_1234 / σt_334
- Fit David's γ σt Improvement: Make dE, θ map
- Fit the π- & proton BCAL shower σt vs θ distributions in dE bins:
- Both the non-smeared and smeared
will apply improvement % to these resolutions.
- Generate, simulate, smear, and reconstruct π(1600) -> b1π events with t-slope = 2.0 (was 5.0 for X(2000))
- For 1234 Segmentation, disabled BCAL hit t-smear in mc_smear, smeared reconstructed shower time as a function of dE & &theta during reconstruction to reach improved resolution.
- Proton, π+'s, π-'s all smeared the same.
- For 1234 Segmentation, disabled BCAL hit t-smear in mc_smear, smeared reconstructed shower time as a function of dE & &theta during reconstruction to reach improved resolution.
David's BCAL Photon Time Resolution Studies
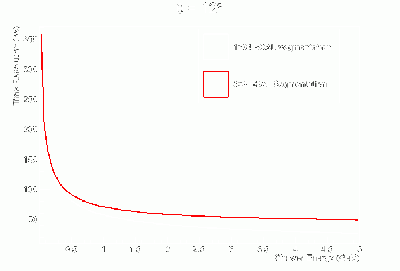
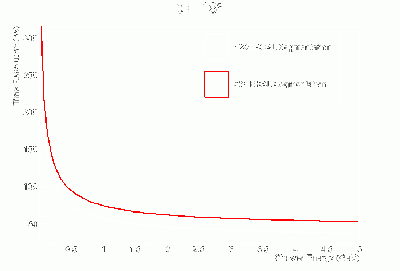
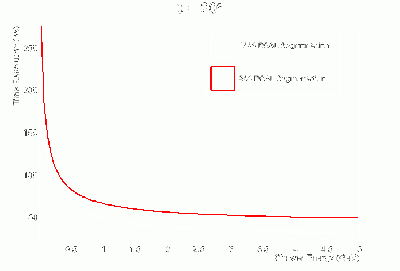
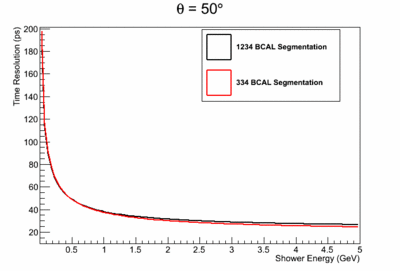
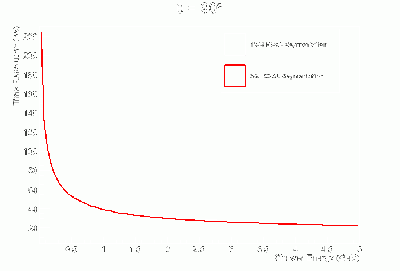
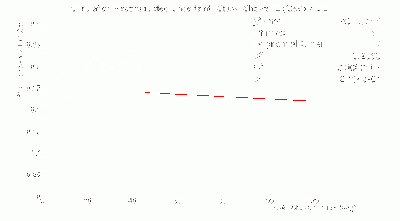
- γ BCAL σt vs. dE (GeV):
Time Resolution Segmentation Ratio
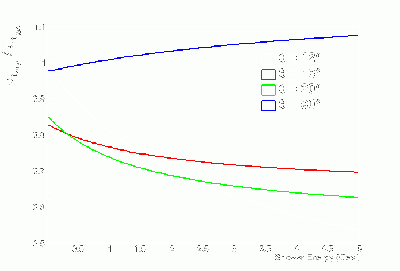
- γ BCAL σt_1234 / σt_334 vs. dE (GeV):
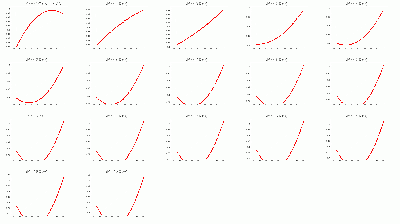
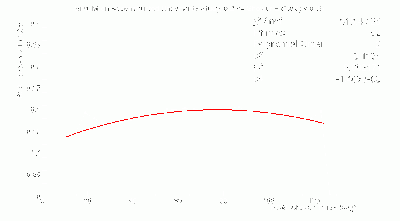
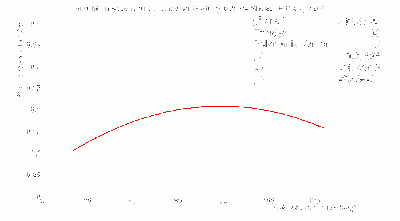
- Fits to γ BCAL σt_1234 / σt_334 vs. θ in dE bins:
- Don't trust fits at low-θ, so linearly-interpolate between 12 and 16 degrees.
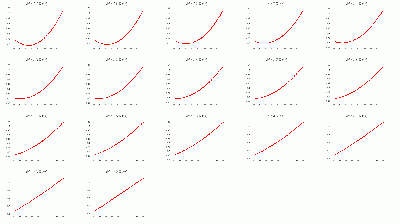
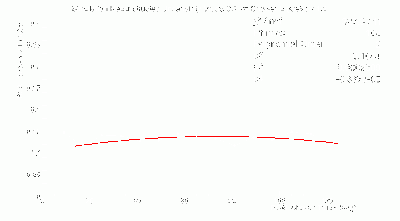
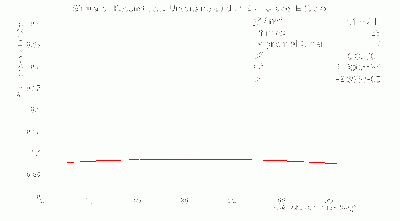
BCAL π- 334 Segmentation σt Fits
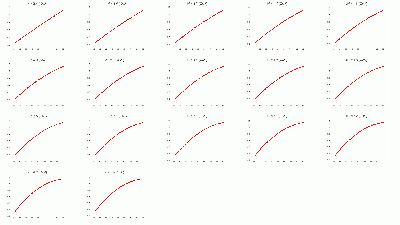
- BCAL π- 334 Segmentation σt Fits vs. θ in dE bins:
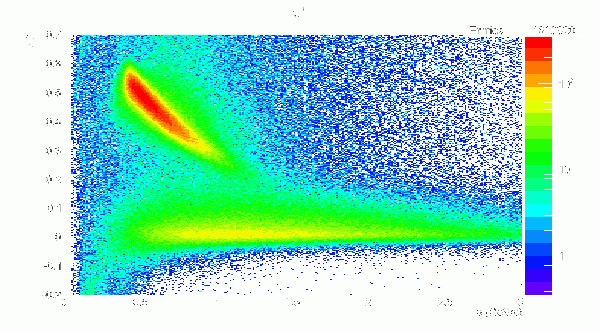
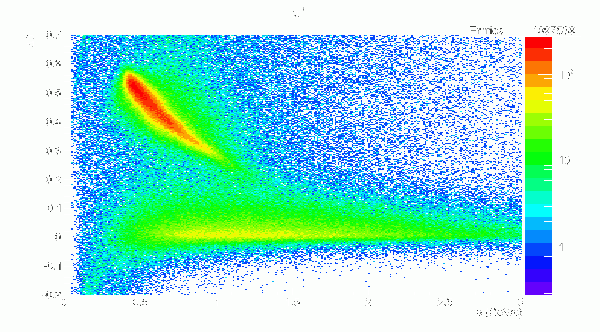
X(2000) -> b1pi Events, π+ BCAL Δβ vs p
Δβ = βp - βt = p/sqrt(p*p + m*m) - path/(c*TOF)
- X(2000) -> b1pi Events, π+ BCAL Δβ vs p, 334 Segmentation:
- X(2000) -> b1pi Events, π+ BCAL Δβ vs p, 1234 Segmentation: